Visualizing Tabular Data + Pandas
Overview
Teaching: 30 min
Exercises: 20 minQuestions
How can I visualize tabular data in Python?
How can I group several plots together?
Objectives
Plot simple graphs from data.
Plot multiple graphs in a single figure.
Visualizing data
The mathematician Richard Hamming once said, “The purpose of computing is insight, not numbers,” and
the best way to develop insight is often to visualize data. Visualization deserves an entire
lecture of its own, but we can explore a few features of Python’s matplotlib library here. While
there is no official plotting library, matplotlib is the de facto standard. First, we will
import the pyplot module from matplotlib and use two of its functions to create and display a
heat map of our data:
import matplotlib.pyplot
image = matplotlib.pyplot.imshow(data[:40,:])
matplotlib.pyplot.show()

Blue pixels in this heat map represent low values, while yellow pixels represent high values. As we can see, the scale of these features varies dramatically, and within a particular feature there is variation between molecules.
ave_feature_value = numpy.mean(data, axis=0)
ave_plot = matplotlib.pyplot.plot(ave_feature_value)
matplotlib.pyplot.show()
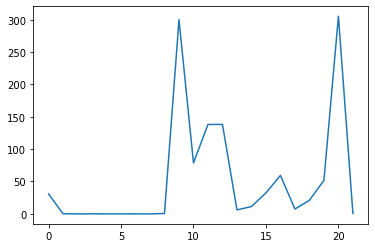
Here, we have put the of each feature across all molecules in the variable
ave_feature_value, then asked matplotlib.pyplot to create and display a line graph of those
values.
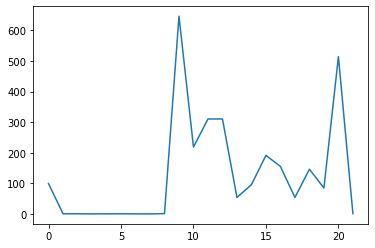
Let’s produce the same for the maximum.
max_plot = matplotlib.pyplot.plot(numpy.max(data, axis=0))
matplotlib.pyplot.show()
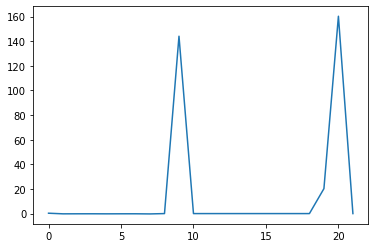
min_plot = matplotlib.pyplot.plot(np.min(data, axis=0))
matplotlib.pyplot.show()
Grouping plots
You can group similar plots in a single figure using subplots.
This script below uses a number of new commands. The function matplotlib.pyplot.figure()
creates a space into which we will place all of our plots. The parameter figsize
tells Python how big to make this space. Each subplot is placed into the figure using
its add_subplot method. The add_subplot method takes 3
parameters. The first denotes how many total rows of subplots there are, the second parameter
refers to the total number of subplot columns, and the final parameter denotes which subplot
your variable is referencing (left-to-right, top-to-bottom). Each subplot is stored in a
different variable (axes1, axes2, axes3). Once a subplot is created, the axes can
be titled using the set_xlabel() command (or set_ylabel()).
Here are our three plots side by side:
import numpy
import matplotlib.pyplot
data = numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',')
array = np.delete(array, 0, 1)
fig = matplotlib.pyplot.figure(figsize=(10.0, 3.0))
axes1 = fig.add_subplot(1, 3, 1)
axes2 = fig.add_subplot(1, 3, 2)
axes3 = fig.add_subplot(1, 3, 3)
axes1.set_ylabel('average')
axes1.plot(numpy.mean(data, axis=0))
axes2.set_ylabel('max')
axes2.plot(numpy.max(data, axis=0))
axes3.set_ylabel('min')
axes3.plot(numpy.min(data, axis=0))
fig.tight_layout()
matplotlib.pyplot.savefig('features.png')
matplotlib.pyplot.show()

The call to loadtxt reads our data,
and the rest of the program tells the plotting library
how large we want the figure to be,
that we’re creating three subplots,
what to draw for each one,
and that we want a tight layout.
(If we leave out that call to fig.tight_layout(),
the graphs will actually be squeezed together more closely.)
The call to savefig stores the plot as a graphics file. This can be
a convenient way to store your plots for use in other documents, web
pages etc. The graphics format is automatically determined by
Matplotlib from the file name ending we specify; here PNG from
‘features.png’. Matplotlib supports many different graphics
formats, including SVG, PDF, and JPEG.
Importing libraries with shortcuts
In this lesson we use the
import matplotlib.pyplotsyntax to import thepyplotmodule ofmatplotlib. However, shortcuts such asimport matplotlib.pyplot as pltare frequently used. Importingpyplotthis way means that after the initial import, rather than writingmatplotlib.pyplot.plot(...), you can now writeplt.plot(...). Another common convention is to use the shortcutimport numpy as npwhen importing the NumPy library. We then can writenp.loadtxt(...)instead ofnumpy.loadtxt(...), for example.Some people prefer these shortcuts as it is quicker to type and results in shorter lines of code - especially for libraries with long names! You will frequently see Python code online using a
pyplotfunction withplt, or a NumPy function withnp, and it’s because they’ve used this shortcut. It makes no difference which approach you choose to take, but you must be consistent as if you useimport matplotlib.pyplot as pltthenmatplotlib.pyplot.plot(...)will not work, and you must useplt.plot(...)instead. Because of this, when working with other people it is important you agree on how libraries are imported.
Plot Scaling
Why do all of our plots stop just short of the upper end of our graph?
Solution
Because matplotlib normally sets x and y axes limits to the min and max of our data (depending on data range)
If we want to change this, we can use the
set_ylim(min, max)method of each ‘axes’, for example:axes3.set_ylim(0,6)Update your plotting code to automatically set a more appropriate scale. (Hint: you can make use of the
maxandminmethods to help.)Solution
# One method axes3.set_ylabel('min') axes3.plot(numpy.min(data, axis=0)) axes3.set_ylim(0,6)Solution
# A more automated approach min_data = numpy.min(data, axis=0) axes3.set_ylabel('min') axes3.plot(min_data) axes3.set_ylim(numpy.min(min_data), numpy.max(min_data) * 1.1)
Drawing Straight Lines
In the center and right subplots above, we expect all lines to look like step functions because non-integer value are not realistic for the minimum and maximum values. However, you can see that the lines are not always vertical or horizontal, and in particular the step function in the subplot on the right looks slanted. Why is this?
Solution
Because matplotlib interpolates (draws a straight line) between the points. One way to do avoid this is to use the Matplotlib
drawstyleoption:import numpy import matplotlib.pyplot data = numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',') fig = matplotlib.pyplot.figure(figsize=(10.0, 3.0)) axes1 = fig.add_subplot(1, 3, 1) axes2 = fig.add_subplot(1, 3, 2) axes3 = fig.add_subplot(1, 3, 3) axes1.set_ylabel('average') axes1.plot(numpy.mean(data, axis=0), drawstyle='steps-mid') axes2.set_ylabel('max') axes2.plot(numpy.max(data, axis=0), drawstyle='steps-mid') axes3.set_ylabel('min') axes3.plot(numpy.min(data, axis=0), drawstyle='steps-mid') fig.tight_layout() matplotlib.pyplot.show()
Make Your Own Plot
Create a plot showing the standard deviation (
numpy.std) of the inflammation data for each day across all patients.Solution
std_plot = matplotlib.pyplot.plot(numpy.std(data, axis=0)) matplotlib.pyplot.show()
Moving Plots Around
Modify the program to display the three plots on top of one another instead of side by side.
Solution
import numpy import matplotlib.pyplot data = numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',') array = np.delete(array, 0, 1) # change figsize (swap width and height) fig = matplotlib.pyplot.figure(figsize=(3.0, 10.0)) # change add_subplot (swap first two parameters) axes1 = fig.add_subplot(3, 1, 1) axes2 = fig.add_subplot(3, 1, 2) axes3 = fig.add_subplot(3, 1, 3) axes1.set_ylabel('average') axes1.plot(numpy.mean(data, axis=0)) axes2.set_ylabel('max') axes2.plot(numpy.max(data, axis=0)) axes3.set_ylabel('min') axes3.plot(numpy.min(data, axis=0)) fig.tight_layout() matplotlib.pyplot.show()
Pandas
Pandas is a powerful dataframe tool that allows you to interact with data in a manner more like an Excel sheet or R - it’s the arrays we have been using in Numpy, but with labeling retained!
import pandas as pd
df = pd.read_csv('Acyclic.csv')
print(df)
Unnamed: 0 %top ER1 ER2 ER3 ER4 ER5 ER6 ER7 EZ ... \
0 0 2.25 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
1 1 2.50 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
2 2 5.00 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
3 3 5.00 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
4 4 3.50 -0.01 -0.17 0.0 -0.01 0.0 0.0 -0.17 1 ...
.. ... ... ... ... ... ... ... ... ... .. ...
174 174 2.50 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
175 175 3.25 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
176 176 5.00 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
177 177 2.00 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
178 178 15.00 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
boron reagent equivs boron reagent \
0 OB(C1=CC=CC=C1)O 2.0
1 OB(C1=CC=CC=C1)O 2.0
2 OB(C1=CC=CC=C1)O 3.0
3 C1(B2OB(C3=CC=CC=C3)OB(C4=CC=CC=C4)O2)=CC=CC=C1 2.0
4 OB(C1=CC=CC=C1)O 2.0
.. ... ...
174 OB(C1=CC=CC=C1)O 2.4
175 CC1=CC(B(O)O)=CC=C1 2.4
176 OB(C1=CC=CC=C1)O 2.0
177 OB(C1=CC=CC=C1)O 2.4
178 OB(C1=CC=CC=C1)O 2.4
ligand mol% Rh mol% ligand solvent \
0 L3.1 1.5 3.0 dioxane/H2O
1 L3.1 1.5 3.3 dioxane/H2O
2 L3.1 2.5 5.5 dioxane/H2O
3 L3.1 1.5 3.0 dioxane/H2O
4 L3.10 1.5 3.6 dioxane
.. ... ... ... ...
174 L3.86 2.5 6.0 dioxane/H2O
175 L3.86 2.5 6.0 dioxane/H2O
176 L3.86 2.5 6.0 dioxane/H2O
177 L3.86 2.5 6.0 dioxane/H2O
178 L3.87 2.5 6.0 dioxane/H2O
substrate temp time yield
0 CC(/C=C/C(C)C)=O 30 3.0 81.0
1 CC(/C=C/CCCCC)=O 30 1.0 84.0
2 O=C(/C=C/C(OC(C)(C)C)=O)OC(C)(C)C 50 3.0 78.0
3 O=C(/C=C/CCC)OC(C)C 50 1.0 73.0
4 CC(/C=C/C(C)C)=O 25 1.0 91.0
.. ... ... ... ...
174 CCCC/C=C/C1=NC2=CC=CC=C2C=C1 80 0.5 56.0
175 CCCC/C=C/C1=NC2=CC=CC=C2C=C1 80 0.5 71.0
176 C12=CC=CC=C1N=CC(/C=C/CCC3=CC=CC=C3)=N2 80 0.5 80.0
177 O=[N+](C1=CC=C(/C=C/CCC2=CC=CC=C2)C=C1)[O-] 80 0.5 35.0
178 O=[N+](C1=CC=C(/C=C/CCC2=CC=CC=C2)C=C1)[O-] 120 0.5 76.0
[179 rows x 40 columns]
This is dramatically different to the array function where we worked with a csv file where the headers had been stripped out, and where we stripped out the first column - the index. Printing this dataframe is very informative, displaying the headers and the indices, plus the size of dataframe. The best thing is we can now do some work using those labels!
There is a lot of data so let’s filter it and remove some columns.
df = df.filter(['solvent', '%top', 'temp', 'time', 'yield'])
print(df)
print(df['solvent']) # print just the solvent column
solvent %top temp time yield
0 dioxane/H2O 2.25 30 3.0 81.0
1 dioxane/H2O 2.50 30 1.0 84.0
2 dioxane/H2O 5.00 50 3.0 78.0
3 dioxane/H2O 5.00 50 1.0 73.0
4 dioxane 3.50 25 1.0 91.0
.. ... ... ... ... ...
174 dioxane/H2O 2.50 80 0.5 56.0
175 dioxane/H2O 3.25 80 0.5 71.0
176 dioxane/H2O 5.00 80 0.5 80.0
177 dioxane/H2O 2.00 80 0.5 35.0
178 dioxane/H2O 15.00 120 0.5 76.0
[179 rows x 5 columns]
0 dioxane/H2O
1 dioxane/H2O
2 dioxane/H2O
3 dioxane/H2O
4 dioxane
...
174 dioxane/H2O
175 dioxane/H2O
176 dioxane/H2O
177 dioxane/H2O
178 dioxane/H2O
Name: solvent, Length: 179, dtype: object
Printing a column now is much more human readable as we just need the name of the column. And when we want to do some work upon that data, we can convert first to a normal array.
df.to_numpy()
array([['dioxane/H2O', 2.25, 30, 3.0, 81.0],
['dioxane/H2O', 2.5, 30, 1.0, 84.0],
['dioxane/H2O', 5.0, 50, 3.0, 78.0],
['dioxane/H2O', 5.0, 50, 1.0, 73.0],
['dioxane', 3.5, 25, 1.0, 91.0],
['MeOH/CH2Cl2', 33.0, 60, 1.0, 89.0],
['MeOH/CH2Cl2', 11.5, 60, 1.0, 99.0],
['MeOH/CH2Cl2', 1.275, 60, 1.0, 92.0],
['i-PrOH/CH2Cl2', 1.833333333, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.666666667, 60, 1.0, 85.0],
['i-PrOH/CH2Cl2', 2.166666667, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.75, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.25, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 2.0, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.0, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.0, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 0.5, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.5, 60, 1.0, 77.0],
['i-PrOH/CH2Cl2', 1.75, 60, 1.0, 99.0],
['MeOH/CH2Cl2', 1.0, 60, 1.0, 83.0],
['i-PrOH/CH2Cl2', 2.5, 60, 1.0, 74.0],
['toluene/H2O', 25.0, 50, 23.0, 67.0],
['dioxane', 11.5, 25, 1.0, 90.0],
['toluene/H2O', 23.0, 50, 23.0, 85.0],
['dioxane', 11.0, 25, 1.0, 72.0],
['dioxane', 8.5, 25, 1.0, 100.0],
['toluene/H2O', 22.5, 50, 23.0, 56.0],
['EtOH', 2.0, 30, 2.0, 89.0],
['EtOH', 2.0, 30, 2.0, 89.0],
['EtOH', 3.5, 30, 8.0, 99.0],
['toluene/H2O', 28.0, 50, 23.0, 98.0],
['dioxane', 4.5, 25, 1.0, 85.0],
['toluene/H2O', 27.5, 50, 23.0, 90.0],
['dioxane', 3.625, 25, 1.0, 87.0],
['EtOH', 1.0, 25, 96.0, 68.0],
['EtOH', 3.25, 25, 11.0, 86.0],
['EtOH', 1.5, 25, 5.5, 72.0],
['EtOH', 2.0, 25, 50.0, 74.0],
['EtOH', 2.5, 25, 25.0, 77.0],
['EtOH', 5.0, 25, 24.0, 94.0],
['EtOH', 1.4, 25, 96.0, 80.0],
['EtOH', 2.75, 25, 16.0, 99.0],
['EtOH', 0.3, 25, 72.0, 99.0],
['EtOH', 11.0, 25, 27.0, 98.0],
['EtOH', 3.0, 25, 16.0, 95.0],
['EtOH', 1.0, 60, 2.0, 81.0],
['EtOH', 0.5, 60, 14.0, 99.5],
['EtOH', 2.0, 60, 23.0, 88.0],
['toluene/H2O', 8.125, 50, 1.5, 85.0],
['dioxane/H2O', 11.0, 30, 1.0, 89.0],
['dioxane/H2O', 4.0, 30, 1.0, 90.0],
['dioxane', 6.0, 25, 1.0, 74.0],
['toluene/H2O', 24.0, 50, 23.0, 95.0],
['dioxane/H2O', 11.0, 30, 1.0, 91.0],
['dioxane/H2O', 7.5, 30, 1.0, 88.0],
['dioxane/H2O', 19.5, 25, 3.0, 95.0],
['MeOH/H2O', 5.0, 50, 1.0, 91.0],
['dioxane/H2O', 5.0, 50, 3.0, 90.0],
['dioxane/H2O', 4.5, 50, 3.0, 80.0],
['dioxane/H2O', 7.0, 50, 3.0, 78.0],
['dioxane/H2O', 5.0, 50, 3.0, 85.0],
['dioxane/H2O', 6.5, 50, 3.0, 91.0],
['dioxane', 1.5, 50, 6.0, 90.0],
['dioxane/H2O', 7.5, 30, 1.0, 95.0],
['dioxane/H2O', 5.75, 30, 1.0, 90.0],
['dioxane/H2O', 1.0, 30, 1.0, 90.0],
['dioxane/H2O', 2.0, 30, 3.0, 93.0],
['dioxane/H2O', 2.5, 30, 1.0, 93.0],
['dioxane', 21.5, 20, 1.0, 99.0],
['dioxane', 1.5, 20, 1.0, 95.0],
['dioxane', 2.0, 20, 1.0, 96.0],
['dioxane', 0.5, 20, 1.0, 94.0],
['dioxane', 1.0, 20, 1.0, 99.0],
['dioxane', 6.25, 20, 1.0, 33.0],
['dioxane', 1.75, 20, 1.0, 95.0],
['dioxane', 1.0, 20, 1.0, 92.0],
['dioxane', 2.0, 20, 1.0, 90.0],
['dioxane', 2.0, 20, 1.0, 92.0],
['dioxane', 1.0, 30, 2.0, 96.0],
['dioxane', 1.0, 30, 2.0, 97.0],
['dioxane', 1.0, 20, 1.0, 91.0],
['dioxane/H2O', 15.0, 50, 20.0, 75.0],
['dioxane', 24.0, 20, 1.0, 74.0],
['dioxane/H2O', 96.0, 25, 3.0, 99.0],
['dioxane/H2O', 91.5, 25, 3.0, 98.0],
['dioxane', 0.5, 50, 6.0, 95.0],
['dioxane', 0.5, 50, 6.0, 94.0],
['dioxane', 3.5, 50, 6.0, 90.0],
['dioxane', 2.5, 50, 6.0, 92.0],
['dioxane', 2.5, 50, 6.0, 94.0],
['dioxane', 2.0, 50, 6.0, 95.0],
['dioxane', 3.5, 50, 6.0, 96.0],
['dioxane', 1.5, 50, 12.0, 91.0],
['dioxane', 1.0, 50, 6.0, 89.0],
['dioxane/H2O', 4.2, 50, 6.0, 94.0],
['dioxane/H2O', 2.5, 50, 6.0, 88.0],
['dioxane/H2O', 95.0, 50, 20.0, 92.0],
['dioxane/H2O', 94.5, 50, 20.0, 83.0],
['dioxane/H2O', 93.5, 50, 20.0, 93.0],
['dioxane/H2O', 96.0, 50, 20.0, 84.0],
['dioxane/H2O', 94.5, 50, 20.0, 83.0],
['dioxane/H2O', 92.5, 50, 20.0, 92.0],
['dioxane/H2O', 96.0, 50, 20.0, 92.0],
['dioxane/H2O', 93.5, 50, 20.0, 86.0],
['dioxane/H2O', 93.0, 50, 20.0, 74.0],
['dioxane/H2O', 90.0, 50, 20.0, 91.0],
['DCM', 16.5, 25, 1.0, 99.0],
['MeOH/CH2Cl2', 24.0, 25, 1.0, 99.0],
['DCM', 2.5, 30, 3.0, 99.0],
['MeOH/CH2Cl2', 1.5, 30, 3.0, 99.0],
['dioxane/H2O', 25.5, 25, 3.0, 99.0],
['dioxane/H2O', 16.5, 25, 3.0, 98.0],
['dioxane/H2O', 21.5, 25, 3.0, 98.0],
['MeOH/CH2Cl2', 12.75, 30, 3.0, 95.0],
['MeOH/CH2Cl2', 13.5, 30, 3.0, 98.0],
['MeOH/CH2Cl2', 7.625, 30, 3.0, 93.0],
['MeOH/CH2Cl2', 12.5, 30, 3.0, 99.0],
['MeOH/CH2Cl2', 14.75, 30, 3.0, 96.0],
['MeOH/CH2Cl2', 6.25, 30, 3.0, 43.0],
['dioxane/H2O', 14.0, 25, 3.0, 98.0],
['dioxane/H2O', 15.0, 80, 16.0, 73.0],
['dioxane/H2O', 7.0, 25, 2.0, 74.0],
['MeOH/CH2Cl2', 95.0, 25, 1.0, 40.0],
['dioxane/H2O', 95.0, 25, 0.1, 68.0],
['dioxane/H2O', 94.5, 25, 1.0, 78.0],
['dioxane/H2O', 94.0, 25, 1.0, 93.0],
['dioxane/H2O', 4.5, 25, 3.0, 99.0],
['dioxane/H2O', 91.5, 50, 1.25, 50.0],
['MeOH/H2O', 96.5, 50, 1.25, 63.0],
['MeOH/H2O', 94.5, 50, 22.0, 70.0],
['MeOH/H2O', 95.5, 50, 2.5, 85.0],
['dioxane/H2O', 96.0, 50, 1.25, 43.0],
['MeOH/H2O', 96.5, 50, 1.25, 90.0],
['MeOH/H2O', 95.5, 50, 4.0, 76.0],
['MeOH/H2O', 96.5, 50, 1.25, 78.0],
['MeOH/H2O', 95.5, 50, 1.25, 87.0],
['MeOH/H2O', 95.0, 50, 1.25, 78.0],
['MeOH/H2O', 95.5, 50, 14.0, 68.0],
['MeOH/H2O', 96.0, 50, 14.0, 62.0],
['dioxane/H2O', 95.0, 50, 14.0, 65.0],
['dioxane/H2O', 96.5, 50, 14.0, 70.0],
['MeOH/H2O', 96.0, 50, 24.0, 69.0],
['MeOH/H2O', 96.0, 50, 1.25, 76.0],
['MeOH/H2O', 96.25, 50, 1.25, 85.0],
['MeOH/H2O', 96.0, 50, 1.25, 78.0],
['MeOH/H2O', 96.0, 50, 1.25, 84.0],
['MeOH/H2O', 97.0, 50, 1.5, 95.0],
['MeOH/H2O', 96.5, 50, 2.0, 93.0],
['MeOH/H2O', 95.5, 50, 18.5, 95.0],
['dioxane/H2O', 74.0, 50, 1.25, 43.0],
['dioxane/H2O', 59.5, 50, 18.0, 71.0],
['dioxane/H2O', 95.5, 50, 18.0, 85.0],
['dioxane/H2O', 80.0, 50, 1.25, 45.0],
['dioxane', 0.3, 50, 3.0, 97.0],
['dioxane/H2O', 3.6, 30, 3.0, 84.0],
['toluene/H2O', 97.5, 100, 12.0, 83.0],
['toluene/H2O', 96.0, 100, 12.0, 72.0],
['toluene/H2O', 97.0, 100, 18.0, 88.0],
['dioxane/H2O', 5.0, 80, 16.0, 41.0],
['toluene', 98.0, 100, 12.0, 82.0],
['dioxane/H2O', 6.5, 80, 0.5, 84.0],
['dioxane/H2O', 2.5, 80, 0.5, 24.0],
['dioxane/H2O', 5.0, 80, 0.5, 44.0],
['dioxane/H2O', 4.5, 80, 0.5, 91.0],
['dioxane/H2O', 2.5, 80, 16.0, 72.0],
['dioxane/H2O', 2.5, 100, 16.0, 62.0],
['dioxane/H2O', 4.0, 100, 16.0, 56.0],
['dioxane/H2O', 3.5, 80, 1.5, 88.0],
['dioxane/H2O', 6.5, 80, 1.0, 79.0],
['dioxane/H2O', 3.0, 80, 0.5, 95.0],
['dioxane/H2O', 1.0, 80, 0.5, 72.0],
['dioxane/H2O', 1.5, 80, 0.5, 65.0],
['dioxane/H2O', 3.5, 80, 0.5, 69.0],
['dioxane/H2O', 3.5, 80, 0.5, 78.0],
['dioxane/H2O', 2.5, 80, 0.5, 56.0],
['dioxane/H2O', 3.25, 80, 0.5, 71.0],
['dioxane/H2O', 5.0, 80, 0.5, 80.0],
['dioxane/H2O', 2.0, 80, 0.5, 35.0],
['dioxane/H2O', 15.0, 120, 0.5, 76.0]], dtype=object)
Key Points
Use the
pyplotmodule from thematplotliblibrary for creating simple visualizations.
