Python Fundamentals
Overview
Teaching: 20 min
Exercises: 10 minQuestions
What basic data types can I work with in Python?
How can I create a new variable in Python?
How do I use a function?
Can I change the value associated with a variable after I create it?
Objectives
Assign values to variables.
Variables
Any Python interpreter can be used as a calculator:
3 + 5 * 4
23
This is great but not very interesting.
To do anything useful with data, we need to assign its value to a variable.
In Python, we can assign a value to a
variable, using the equals sign =.
For example, we can track the weight of a patient who weighs 60 kilograms by
assigning the value 60 to a variable weight_kg:
weight_kg = 60
From now on, whenever we use weight_kg, Python will substitute the value we assigned to
it. In layman’s terms, a variable is a name for a value.
In Python, variable names:
- can include letters, digits, and underscores
- cannot start with a digit
- are case sensitive.
This means that, for example:
weight0is a valid variable name, whereas0weightis notweightandWeightare different variables
Types of data
Python knows various types of data. Three common ones are:
- integer numbers
- floating point numbers, and
- strings.
In the example above, variable weight_kg has an integer value of 60.
If we want to more precisely track the weight of our patient,
we can use a floating point value by executing:
weight_kg = 60.3
To create a string, we add single or double quotes around some text. To identify and track a patient throughout our study, we can assign each person a unique identifier by storing it in a string:
patient_id = '001'
Using Variables in Python
Once we have data stored with variable names, we can make use of it in calculations. We may want to store our patient’s weight in pounds as well as kilograms:
weight_lb = 2.2 * weight_kg
We might decide to add a prefix to our patient identifier:
patient_id = 'inflam_' + patient_id
Built-in Python functions
To carry out common tasks with data and variables in Python,
the language provides us with several built-in functions.
To display information to the screen, we use the print function:
print(weight_lb)
print(patient_id)
132.66
inflam_001
When we want to make use of a function, referred to as calling the function,
we follow its name by parentheses. The parentheses are important:
if you leave them off, the function doesn’t actually run!
Sometimes you will include values or variables inside the parentheses for the function to use.
In the case of print,
we use the parentheses to tell the function what value we want to display.
We will learn more about how functions work and how to create our own in later episodes.
We can display multiple things at once using only one print call:
print(patient_id, 'weight in kilograms:', weight_kg)
inflam_001 weight in kilograms: 60.3
We can also call a function inside of another
function call.
For example, Python has a built-in function called type that tells you a value’s data type:
print(type(60.3))
print(type(patient_id))
<class 'float'>
<class 'str'>
Moreover, we can do arithmetic with variables right inside the print function:
print('weight in pounds:', 2.2 * weight_kg)
weight in pounds: 132.66
The above command, however, did not change the value of weight_kg:
print(weight_kg)
60.3
To change the value of the weight_kg variable, we have to
assign weight_kg a new value using the equals = sign:
weight_kg = 65.0
print('weight in kilograms is now:', weight_kg)
weight in kilograms is now: 65.0
Variables as Sticky Notes
A variable in Python is analogous to a sticky note with a name written on it: assigning a value to a variable is like putting that sticky note on a particular value.
Using this analogy, we can investigate how assigning a value to one variable does not change values of other, seemingly related, variables. For example, let’s store the subject’s weight in pounds in its own variable:
# There are 2.2 pounds per kilogram weight_lb = 2.2 * weight_kg print('weight in kilograms:', weight_kg, 'and in pounds:', weight_lb)weight in kilograms: 65.0 and in pounds: 143.0
Similar to above, the expression
2.2 * weight_kgis evaluated to143.0, and then this value is assigned to the variableweight_lb(i.e. the sticky noteweight_lbis placed on143.0). At this point, each variable is “stuck” to completely distinct and unrelated values.Let’s now change
weight_kg:weight_kg = 100.0 print('weight in kilograms is now:', weight_kg, 'and weight in pounds is still:', weight_lb)weight in kilograms is now: 100.0 and weight in pounds is still: 143.0
Since
weight_lbdoesn’t “remember” where its value comes from, it is not updated when we changeweight_kg.
Check Your Understanding
What values do the variables
massandagehave after each of the following statements? Test your answer by executing the lines.mass = 47.5 age = 122 mass = mass * 2.0 age = age - 20Solution
`mass` holds a value of 47.5, `age` does not exist `mass` still holds a value of 47.5, `age` holds a value of 122 `mass` now has a value of 95.0, `age`'s value is still 122 `mass` still has a value of 95.0, `age` now holds 102
Sorting Out References
Python allows you to assign multiple values to multiple variables in one line by separating the variables and values with commas. What does the following program print out?
first, second = 'Grace', 'Hopper' third, fourth = second, first print(third, fourth)Solution
Hopper Grace
Seeing Data Types
What are the data types of the following variables?
planet = 'Earth' apples = 5 distance = 10.5Solution
type(planet) type(apples) type(distance)<class 'str'> <class 'int'> <class 'float'>
Key Points
Basic data types in Python include integers, strings, and floating-point numbers.
Use
variable = valueto assign a value to a variable in order to record it in memory.Variables are created on demand whenever a value is assigned to them.
Use
print(something)to display the value ofsomething.Built-in functions are always available to use.
Analyzing Molecular Data
Overview
Teaching: 40 min
Exercises: 20 minQuestions
How can I process tabular data files in Python?
Objectives
Explain what a library is and what libraries are used for.
Import a Python library and use the functions it contains.
Read tabular data from a file into a program.
Select individual values and subsections from data.
Perform operations on arrays of data.
Words are useful, but what’s more useful are the sentences and stories we build with them. Similarly, while a lot of powerful, general tools are built into Python, specialized tools built up from these basic units live in libraries that can be called upon when needed.
Loading data into Python
To begin processing inflammation data, we need to load it into Python. We can do that using a library called NumPy, which stands for Numerical Python. In general, you should use this library when you want to do fancy things with lots of numbers, especially if you have matrices or arrays. To tell Python that we’d like to start using NumPy, we need to import it:
import numpy
Importing a library is like getting a piece of lab equipment out of a storage locker and setting it up on the bench. Libraries provide additional functionality to the basic Python package, much like a new piece of equipment adds functionality to a lab space. Just like in the lab, importing too many libraries can sometimes complicate and slow down your programs - so we only import what we need for each program.
Once we’ve imported the library, we can ask the library to read our data file for us:
numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',')
array([[ 0.00000000e+00 6.00000000e+00 -9.00000000e-02 ... 2.02559852e+01
2.79454641e+02 1.00000000e+00]
[ 1.00000000e+00 2.33333333e+00 -9.00000000e-02 ... 2.02559852e+01
2.79454641e+02 1.00000000e+00]
[ 2.00000000e+00 7.90000000e+01 -1.40000000e-01 ... 7.60741678e+01
2.23068184e+02 5.00000000e-01]
...
[ 3.40000000e+02 2.00000000e+00 3.60000000e-01 ... 8.45799261e+01
3.94431707e+02 5.00000000e-01]
[ 3.41000000e+02 1.50000000e+01 3.60000000e-01 ... 8.45799261e+01
2.82386669e+02 5.00000000e-01]
[ 3.42000000e+02 3.00000000e-01 0.00000000e+00 ... 8.45799261e+01
2.43000629e+02 5.00000000e-01]])
The expression numpy.loadtxt(...) is a
function call
that asks Python to run the function loadtxt which
belongs to the numpy library.
This dotted notation
is used everywhere in Python: the thing that appears before the dot contains the thing that
appears after.
As an example, John Smith is the John that belongs to the Smith family.
We could use the dot notation to write his name smith.john,
just as loadtxt is a function that belongs to the numpy library.
numpy.loadtxt has two parameters: the name of the file
we want to read and the delimiter that separates values
on a line. These both need to be character strings
(or strings for short), so we put them in quotes.
Since we haven’t told it to do anything else with the function’s output,
the notebook displays it.
In this case,
that output is the data we just loaded.
By default,
only a few rows and columns are shown
(with ... to omit elements when displaying big arrays).
Note that, to save space when displaying NumPy arrays, Python does not show us trailing zeros,
so 1.0 becomes 1..
Our call to numpy.loadtxt read our file
but didn’t save the data in memory.
To do that,
we need to assign the array to a variable. In a similar manner to how we assign a single
value to a variable, we can also assign an array of values to a variable using the same syntax.
Let’s re-run numpy.loadtxt and save the returned data:
data = numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',')
This statement doesn’t produce any output because we’ve assigned the output to the variable data.
If we want to check that the data have been loaded,
we can print the variable’s value:
print(data)
[[ 0.00000000e+00 6.00000000e+00 -9.00000000e-02 ... 2.02559852e+01
2.79454641e+02 1.00000000e+00]
[ 1.00000000e+00 2.33333333e+00 -9.00000000e-02 ... 2.02559852e+01
2.79454641e+02 1.00000000e+00]
[ 2.00000000e+00 7.90000000e+01 -1.40000000e-01 ... 7.60741678e+01
2.23068184e+02 5.00000000e-01]
...
[ 3.40000000e+02 2.00000000e+00 3.60000000e-01 ... 8.45799261e+01
3.94431707e+02 5.00000000e-01]
[ 3.41000000e+02 1.50000000e+01 3.60000000e-01 ... 8.45799261e+01
2.82386669e+02 5.00000000e-01]
[ 3.42000000e+02 3.00000000e-01 0.00000000e+00 ... 8.45799261e+01
2.43000629e+02 5.00000000e-01]]
Now that the data are in memory,
we can manipulate them.
First,
let’s ask what type of thing data refers to:
print(type(data))
<class 'numpy.ndarray'>
The output tells us that data currently refers to
an N-dimensional array, the functionality for which is provided by the NumPy library.
These data correspond to arthritis patients’ inflammation.
The rows are the individual patients, and the columns
are their daily inflammation measurements.
Data Type
A Numpy array contains one or more elements of the same type. The
typefunction will only tell you that a variable is a NumPy array but won’t tell you the type of thing inside the array. We can find out the type of the data contained in the NumPy array.print(data.dtype)float64This tells us that the NumPy array’s elements are floating-point numbers.
With the following command, we can see the array’s shape:
print(data.shape)
(343, 23)
The output tells us that the data array variable contains 343 rows and 23 columns. When we
created the variable data to store our arthritis data, we did not only create the array; we also
created information about the array, called members or
attributes. This extra information describes data in the same way an adjective describes a noun.
data.shape is an attribute of data which describes the dimensions of data. We use the same
dotted notation for the attributes of variables that we use for the functions in libraries because
they have the same part-and-whole relationship.
If we want to get a single number from the array, we must provide an index in square brackets after the variable name, just as we do in math when referring to an element of a matrix. Our molecular properties data has two dimensions, so we will need to use two indices to refer to one specific value:
print('first value in data:', data[0, 0])
first value in data: 0.0
print('middle value in data:', data[172, 11])
middle value in data: 94.18429025
The expression data[172, 11] accesses the element at row 172, column 11. While this expression may
not surprise you,
data[0, 0] might.
Programming languages like Fortran, MATLAB and R start counting at 1
because that’s what human beings have done for thousands of years.
Languages in the C family (including C++, Java, Perl, and Python) count from 0
because it represents an offset from the first value in the array (the second
value is offset by one index from the first value). This is closer to the way
that computers represent arrays (if you are interested in the historical
reasons behind counting indices from zero, you can read
Mike Hoye’s blog post).
As a result,
if we have an M×N array in Python,
its indices go from 0 to M-1 on the first axis
and 0 to N-1 on the second.
It takes a bit of getting used to,
but one way to remember the rule is that
the index is how many steps we have to take from the start to get the item we want.
In the Corner
What may also surprise you is that when Python displays an array, it shows the element with index
[0, 0]in the upper left corner rather than the lower left. This is consistent with the way mathematicians draw matrices but different from the Cartesian coordinates. The indices are (row, column) instead of (column, row) for the same reason, which can be confusing when plotting data.
Slicing data
An index like [30, 20] selects a single element of an array,
but we can select whole sections as well.
For example,
we can select the first properties (columns) of values
for the first four molecules (rows) like this:
print(data[0:4, 0:10])
[[ 0.00000000e+00 6.00000000e+00 -9.00000000e-02 0.00000000e+00
0.00000000e+00 -9.00000000e-02 0.00000000e+00 0.00000000e+00
0.00000000e+00 0.00000000e+00]
[ 1.00000000e+00 2.33333333e+00 -9.00000000e-02 0.00000000e+00
0.00000000e+00 -9.00000000e-02 0.00000000e+00 0.00000000e+00
0.00000000e+00 0.00000000e+00]
[ 2.00000000e+00 7.90000000e+01 -1.40000000e-01 0.00000000e+00
0.00000000e+00 0.00000000e+00 -1.70000000e-01 0.00000000e+00
-2.70000000e-01 0.00000000e+00]
[ 3.00000000e+00 9.75000000e+01 7.00000000e-02 0.00000000e+00
0.00000000e+00 -1.20000000e-01 -1.70000000e-01 0.00000000e+00
-2.70000000e-01 0.00000000e+00]]
The slice 0:4 means, “Start at index 0 and go up to,
but not including, index 4”. Again, the up-to-but-not-including takes a bit of getting used to,
but the rule is that the difference between the upper and lower bounds is the number of values in
the slice.
We don’t have to start slices at 0:
print(data[5:10, 0:10])
[[ 5.000e+00 9.575e+01 0.000e+00 0.000e+00 0.000e+00 0.000e+00
-1.700e-01 0.000e+00 -2.700e-01 0.000e+00]
[ 6.000e+00 9.650e+01 -1.600e-01 0.000e+00 0.000e+00 -1.000e-02
-1.700e-01 0.000e+00 -2.700e-01 0.000e+00]
[ 7.000e+00 9.400e+01 -9.000e-02 0.000e+00 0.000e+00 -1.000e-02
-1.700e-01 0.000e+00 -2.700e-01 0.000e+00]
[ 8.000e+00 1.250e+00 0.000e+00 0.000e+00 0.000e+00 -1.700e-01
0.000e+00 0.000e+00 -1.500e-01 0.000e+00]
[ 9.000e+00 2.250e+00 -9.000e-02 0.000e+00 0.000e+00 -9.000e-02
0.000e+00 0.000e+00 0.000e+00 1.000e+00]]
We also don’t have to include the upper and lower bound on the slice. If we don’t include the lower bound, Python uses 0 by default; if we don’t include the upper, the slice runs to the end of the axis, and if we don’t include either (i.e., if we use ‘:’ on its own), the slice includes everything:
small = data[:3, 18:]
print('small is:')
print(small)
The above example selects rows 0 through 2 and columns 36 through to the end of the array.
small is:
[[ 0. 0. 20.25598522 279.4546413 1. ]
[ 0. 0. 20.25598522 279.4546413 1. ]
[ 21.5742147 0. 76.07416785 223.0681843 0.5 ]]
Analyzing data
NumPy has several useful functions that take an array as input to perform operations on its values.
If we want to find the average inflammation for all patients on
all days, for example, we can ask NumPy to compute data’s mean value:
Let’s first remove the first column, our data lables which have been interpreted as real numbers
data = np.delete(data, 0, 1)
And now we can compute the mean
print(numpy.mean(data))
53.72029602831553
mean is a function that takes
an array as an argument.
Not All Functions Have Input
Generally, a function uses inputs to produce outputs. However, some functions produce outputs without needing any input. For example, checking the current time doesn’t require any input.
import time print(time.ctime())Sat Mar 26 13:07:33 2016For functions that don’t take in any arguments, we still need parentheses (
()) to tell Python to go and do something for us.
Let’s use three other NumPy functions to get some descriptive values about the dataset. We’ll also use multiple assignment, a convenient Python feature that will enable us to do this all in one line.
maxval, minval, stdval = numpy.max(data), numpy.min(data), numpy.std(data)
print('maximum property:', maxval)
print('minimum property:', minval)
print('standard deviation:', stdval)
Here we’ve assigned the return value from numpy.max(data) to the variable maxval, the value
from numpy.min(data) to minval, and so on.
maximum property: 646.1013477
minimum property: -0.27
standard deviation: 98.0360663022302
Mystery Functions in IPython
How did we know what functions NumPy has and how to use them? If you are working in IPython or in a Jupyter Notebook, there is an easy way to find out. If you type the name of something followed by a dot, then you can use tab completion (e.g. type
numpy.and then press Tab) to see a list of all functions and attributes that you can use. After selecting one, you can also add a question mark (e.g.numpy.cumprod?), and IPython will return an explanation of the method! This is the same as doinghelp(numpy.cumprod). Similarly, if you are using the “plain vanilla” Python interpreter, you can typenumpy.and press the Tab key twice for a listing of what is available. You can then use thehelp()function to see an explanation of the function you’re interested in, for example:help(numpy.cumprod).
When analyzing data, though, we often want to look at variations in statistical values, such as the maximum feature per molcule or the average of a feature across all molecules. One way to do this is to create a new temporary array of the data we want, then ask it to do the calculation:
molecule_0 = data[0, :] # 0 on the first axis (rows), everything on the second (columns)
print('maximum property for molecule 0:', numpy.max(patient_0))
maximum property for molecule 0: 279.4546413
Everything in a line of code following the ‘#’ symbol is a comment that is ignored by Python. Comments allow programmers to leave explanatory notes for other programmers or their future selves.
We don’t actually need to store the row in a variable of its own. Instead, we can combine the selection and the function call:
print('maximum feature for molecule 2:', numpy.max(data[2, :]))
maximum feature value for molecule 2: 223.0681843
What if we need the maximum feature value for each molecule (as in the next diagram on the left) or the average for each feature over all molecules(as in the diagram on the right)? As the diagram below shows, we want to perform the operation across an axis:
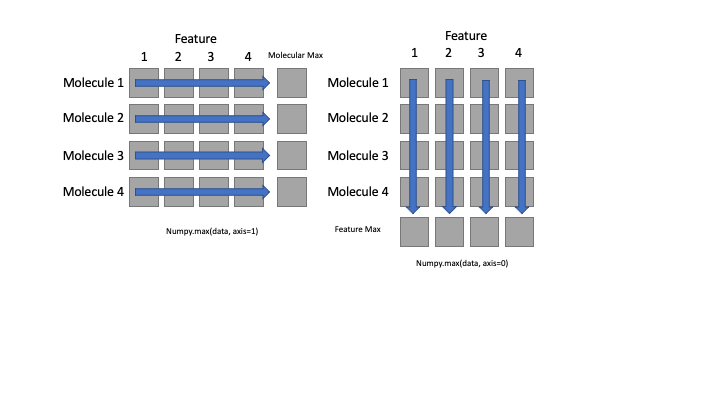
To support this functionality, most array functions allow us to specify the axis we want to work on. If we ask for the average across axis 0 (rows in our 2D example), we get:
print(numpy.mean(data, axis=0))
[ 3.08195303e+01 2.47230321e-02 -2.65597668e-02 -1.11370262e-02
-2.74344023e-02 -4.00874636e-02 -1.54810496e-02 -1.09766764e-01
5.18950437e-01 3.00224099e+02 7.87582274e+01 1.38170875e+02
1.38170875e+02 6.06081019e+00 1.13154699e+01 3.23403644e+01
5.94126474e+01 7.38168047e+00 2.10248945e+01 5.16751691e+01
3.05429392e+02 7.49271137e-01]
As a quick check, we can ask this array what its shape is:
print(numpy.mean(data, axis=0).shape)
(22,)
The expression (22,) tells us we have an N×1 vector,
so this is the average for each feature for all patients.
If we average across axis 1 (columns in our 2D example), we get:
print(numpy.mean(data, axis=1))
[51.99058494 51.82391827 34.7444164 47.51821009 53.10204261 46.88745401
56.56150193 62.33442536 36.76296138 51.86558494 51.93376676 51.87694857
51.99058494 51.96785766 51.99058494 52.39967585 48.50888887 51.17339899
52.51430808 51.53703536 51.07226263 51.09764142 51.09006566 51.11279293
51.09385354 51.07112626 51.10521717 51.05976263 51.05976263 51.03703536
51.0824899 51.09385354 51.01430808 51.12794445 52.15067172 59.09881634
59.62154362 81.7668866 84.71718103 85.35354466 64.65989297 70.35633877
64.69777176 47.56366464 36.79705229 43.97138102 46.05687761 53.92547678
65.19569877 58.29873336 61.52907595 25.90992692 38.79831223 47.22487446
47.56474037 29.25618039 36.74523411 33.06116179 40.50612461 35.00590436
64.65231721 64.65231721 64.72049903 64.62958994 64.76595358 64.60686267
64.77731721 65.83413539 55.04347299 56.08892754 70.52307229 70.40375411
70.50602684 70.42648138 70.44920865 70.47193593 70.58557229 70.42193593
70.48329956 70.37193593 70.85829956 70.4946632 70.40375411 70.38102684
70.44920865 70.72761774 37.86605323 54.08456769 40.9980353 65.28660787
42.40487934 67.48111633 68.29929815 40.97894439 58.73055154 49.87170881
61.48362141 54.46929089 45.68393372 42.80692601 48.66797978 45.77484281
43.97138102 48.75888887 48.69070705 74.68138246 74.65865519 74.77229156
74.68138246 74.74956428 78.61320065 78.65865519 78.77229156 78.54501883
78.59047337 78.61320065 78.59047337 46.09096852 46.00005943 46.06824125
45.69011424 45.71238697 39.69894439 46.0227867 46.06824125 46.04551397
46.36369579 46.15915034 46.28415034 46.06824125 46.11369579 46.13642306
46.0227867 46.00005943 46.00005943 46.00005943 46.0227867 46.06824125
46.03415034 46.0227867 46.0227867 50.47733215 46.09096852 47.00005943
46.09096852 46.11369579 46.04551397 46.06824125 46.30687761 46.10233215
46.06824125 46.11369579 46.0227867 46.11369579 46.06824125 46.06824125
46.06824125 46.70460488 46.00005943 46.0227867 46.06824125 46.04551397
46.00005943 46.0227867 46.04551397 46.00005943 46.00005943 46.68187761
47.11369579 73.67545405 86.82517217 53.60184042 53.60184042 53.73820405
53.69274951 53.69274951 53.67002223 53.73820405 53.64729496 53.62456769
53.85184042 53.77002223 53.82911314 53.69274951 54.32911314 53.71547678
49.2855413 49.26281403 49.21735948 49.33099585 49.26281403 49.17190494
49.33099585 49.21735948 49.19463221 49.05826857 50.79240942 51.5651367
51.90604579 50.8151367 50.8151367 50.88331851 50.79240942 50.79240942
50.79240942 63.03017427 89.04005178 89.10823359 44.67502991 89.06277905
89.13096087 88.62524944 23.94910859 54.63289747 59.27904479 61.31193575
68.90962583 80.16916601 73.51560969 76.52277046 99.19594174 98.01204849
27.78408999 50.54895434 25.83445497 48.76477387 48.50888887 40.86076257
49.68989063 46.25807245 50.52622706 50.79240942 88.99459723 50.59440888
50.82168161 51.17339899 42.71601692 64.66241822 43.9941083 43.92592648
43.7441083 43.90319921 47.86825334 47.27734425 47.61825334 25.79629055
47.56366464 57.69435517 47.45002827 47.427301 47.40457373 47.427301
56.77609633 56.57155087 47.54093736 36.75159775 47.56366464 38.23657776
38.21385049 44.03956284 38.21385049 38.32748685 42.30475958 42.19112321
38.66194859 36.72432502 53.33762084 53.24671175 53.2921663 53.31489357
53.33762084 53.2921663 53.33762084 53.2921663 53.26943903 53.2921663
53.31489357 53.26943903 53.33762084 53.31489357 53.31489357 53.32625721
53.31489357 53.31489357 53.36034812 53.33762084 53.2921663 42.87871884
42.21962793 43.85599157 53.96748921 51.74058494 36.76523411 37.29705229
44.00547193 67.5038436 36.74705229 38.19112321 51.83149403 64.60686267
51.78603948 51.87694857 51.80876676 36.77432502 44.03956284 36.91523411
36.75159775 41.18341593 41.11523411 41.16068865 36.75159775 51.78603948
43.92592648 36.77432502 43.51683557 36.81977956 51.80876676 46.92941991
48.59979796 57.51253699 61.73980971 51.80876676 36.71977956 54.30136226
51.78129472 51.98198261 72.39885502 72.30794593 72.30794593 72.37612775
72.35340048 72.48976411 72.33067321 71.62361426 71.64634153 71.73725062
71.73725062 71.69179607 71.72588698 71.80543244 71.6690688 47.11114202
36.71977956]
which is the average of all features per molecule.
Slicing Strings
A section of an array is called a slice. We can take slices of character strings as well:
element = 'oxygen' print('first three characters:', element[0:3]) print('last three characters:', element[3:6])first three characters: oxy last three characters: genWhat is the value of
element[:4]? What aboutelement[4:]? Orelement[:]?Solution
oxyg en oxygenWhat is
element[-1]? What iselement[-2]?Solution
n eGiven those answers, explain what
element[1:-1]does.Solution
Creates a substring from index 1 up to (not including) the final index, effectively removing the first and last letters from ‘oxygen’
How can we rewrite the slice for getting the last three characters of
element, so that it works even if we assign a different string toelement? Test your solution with the following strings:carpentry,clone,hi.Solution
element = 'oxygen' print('last three characters:', element[-3:]) element = 'carpentry' print('last three characters:', element[-3:]) element = 'clone' print('last three characters:', element[-3:]) element = 'hi' print('last three characters:', element[-3:])last three characters: gen last three characters: try last three characters: one last three characters: hi
Thin Slices
The expression
element[3:3]produces an empty string, i.e., a string that contains no characters. Ifdataholds our array of patient data, what doesdata[3:3, 4:4]produce? What aboutdata[3:3, :]?Solution
array([], shape=(0, 0), dtype=float64) array([], shape=(0, 22), dtype=float64)
Stacking Arrays
Arrays can be concatenated and stacked on top of one another, using NumPy’s
vstackandhstackfunctions for vertical and horizontal stacking, respectively.import numpy A = numpy.array([[1,2,3], [4,5,6], [7, 8, 9]]) print('A = ') print(A) B = numpy.hstack([A, A]) print('B = ') print(B) C = numpy.vstack([A, A]) print('C = ') print(C)A = [[1 2 3] [4 5 6] [7 8 9]] B = [[1 2 3 1 2 3] [4 5 6 4 5 6] [7 8 9 7 8 9]] C = [[1 2 3] [4 5 6] [7 8 9] [1 2 3] [4 5 6] [7 8 9]]Write some additional code that slices the first and last columns of
A, and stacks them into a 3x2 array. Make sure toSolution
A ‘gotcha’ with array indexing is that singleton dimensions are dropped by default. That means
A[:, 0]is a one dimensional array, which won’t stack as desired. To preserve singleton dimensions, the index itself can be a slice or array. For example,A[:, :1]returns a two dimensional array with one singleton dimension (i.e. a column vector).D = numpy.hstack((A[:, :1], A[:, -1:])) print('D = ') print(D)D = [[1 3] [4 6] [7 9]]Solution
An alternative way to achieve the same result is to use Numpy’s delete function to remove the second column of A.
D = numpy.delete(A, 1, 1) print('D = ') print(D)D = [[1 3] [4 6] [7 9]]
Change In Values
Let’s presume for a moment that the data for each molecule is logitudinal in nature. So each value is part of a series over time. We might be interested in the change in that value over time. Let’s find out how to calculate changes in the data contained in an array with NumPy.
The
numpy.diff()function takes an array and returns the differences between two successive values. Let’s use it to examine the changes each day across the first week of patient 3 from our inflammation dataset.molecule3_first7features = data[3, :7] print(molecule3_first7features)[79. -0.14 0. 0. 0. -0.17 0. ]Calling
numpy.diff(molecule3_first7features)would do the following calculations[-0.14 - 79, 0 - -0.14, 0 - 0, 0 - 0, -0.17 - 0, 0 - -0.17]and return the 6 difference values in a new array.
numpy.diff(molecule3_first7features)array([-79.14 0.14 0. 0. -0.17 0.17])Note that the array of differences is shorter by one element (length 6).
Key Points
Import a library into a program using
import libraryname.Use the
numpyandpandaslibrary to work with arrays in Python.The expression
array.shapegives the shape of an array.Use
array[x, y]to select a single element from a 2D array.Array indices start at 0, not 1.
Use
low:highto specify aslicethat includes the indices fromlowtohigh-1.Use
# some kind of explanationto add comments to programs.Use
numpy.mean(array),numpy.max(array), andnumpy.min(array)to calculate simple statistics.Use
numpy.mean(array, axis=0)ornumpy.mean(array, axis=1)to calculate statistics across the specified axis.
Visualizing Tabular Data + Pandas
Overview
Teaching: 30 min
Exercises: 20 minQuestions
How can I visualize tabular data in Python?
How can I group several plots together?
Objectives
Plot simple graphs from data.
Plot multiple graphs in a single figure.
Visualizing data
The mathematician Richard Hamming once said, “The purpose of computing is insight, not numbers,” and
the best way to develop insight is often to visualize data. Visualization deserves an entire
lecture of its own, but we can explore a few features of Python’s matplotlib library here. While
there is no official plotting library, matplotlib is the de facto standard. First, we will
import the pyplot module from matplotlib and use two of its functions to create and display a
heat map of our data:
import matplotlib.pyplot
image = matplotlib.pyplot.imshow(data[:40,:])
matplotlib.pyplot.show()

Blue pixels in this heat map represent low values, while yellow pixels represent high values. As we can see, the scale of these features varies dramatically, and within a particular feature there is variation between molecules.
ave_feature_value = numpy.mean(data, axis=0)
ave_plot = matplotlib.pyplot.plot(ave_feature_value)
matplotlib.pyplot.show()
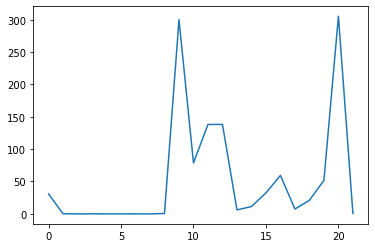
Here, we have put the of each feature across all molecules in the variable
ave_feature_value, then asked matplotlib.pyplot to create and display a line graph of those
values.
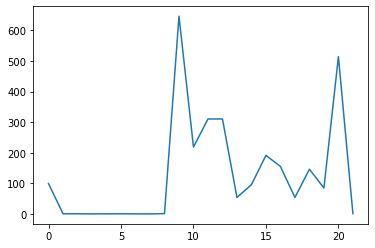
Let’s produce the same for the maximum.
max_plot = matplotlib.pyplot.plot(numpy.max(data, axis=0))
matplotlib.pyplot.show()
min_plot = matplotlib.pyplot.plot(np.min(data, axis=0))
matplotlib.pyplot.show()
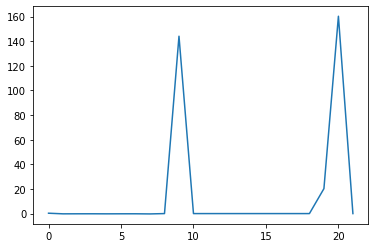
Grouping plots
You can group similar plots in a single figure using subplots.
This script below uses a number of new commands. The function matplotlib.pyplot.figure()
creates a space into which we will place all of our plots. The parameter figsize
tells Python how big to make this space. Each subplot is placed into the figure using
its add_subplot method. The add_subplot method takes 3
parameters. The first denotes how many total rows of subplots there are, the second parameter
refers to the total number of subplot columns, and the final parameter denotes which subplot
your variable is referencing (left-to-right, top-to-bottom). Each subplot is stored in a
different variable (axes1, axes2, axes3). Once a subplot is created, the axes can
be titled using the set_xlabel() command (or set_ylabel()).
Here are our three plots side by side:
import numpy
import matplotlib.pyplot
data = numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',')
array = np.delete(array, 0, 1)
fig = matplotlib.pyplot.figure(figsize=(10.0, 3.0))
axes1 = fig.add_subplot(1, 3, 1)
axes2 = fig.add_subplot(1, 3, 2)
axes3 = fig.add_subplot(1, 3, 3)
axes1.set_ylabel('average')
axes1.plot(numpy.mean(data, axis=0))
axes2.set_ylabel('max')
axes2.plot(numpy.max(data, axis=0))
axes3.set_ylabel('min')
axes3.plot(numpy.min(data, axis=0))
fig.tight_layout()
matplotlib.pyplot.savefig('features.png')
matplotlib.pyplot.show()

The call to loadtxt reads our data,
and the rest of the program tells the plotting library
how large we want the figure to be,
that we’re creating three subplots,
what to draw for each one,
and that we want a tight layout.
(If we leave out that call to fig.tight_layout(),
the graphs will actually be squeezed together more closely.)
The call to savefig stores the plot as a graphics file. This can be
a convenient way to store your plots for use in other documents, web
pages etc. The graphics format is automatically determined by
Matplotlib from the file name ending we specify; here PNG from
‘features.png’. Matplotlib supports many different graphics
formats, including SVG, PDF, and JPEG.
Importing libraries with shortcuts
In this lesson we use the
import matplotlib.pyplotsyntax to import thepyplotmodule ofmatplotlib. However, shortcuts such asimport matplotlib.pyplot as pltare frequently used. Importingpyplotthis way means that after the initial import, rather than writingmatplotlib.pyplot.plot(...), you can now writeplt.plot(...). Another common convention is to use the shortcutimport numpy as npwhen importing the NumPy library. We then can writenp.loadtxt(...)instead ofnumpy.loadtxt(...), for example.Some people prefer these shortcuts as it is quicker to type and results in shorter lines of code - especially for libraries with long names! You will frequently see Python code online using a
pyplotfunction withplt, or a NumPy function withnp, and it’s because they’ve used this shortcut. It makes no difference which approach you choose to take, but you must be consistent as if you useimport matplotlib.pyplot as pltthenmatplotlib.pyplot.plot(...)will not work, and you must useplt.plot(...)instead. Because of this, when working with other people it is important you agree on how libraries are imported.
Plot Scaling
Why do all of our plots stop just short of the upper end of our graph?
Solution
Because matplotlib normally sets x and y axes limits to the min and max of our data (depending on data range)
If we want to change this, we can use the
set_ylim(min, max)method of each ‘axes’, for example:axes3.set_ylim(0,6)Update your plotting code to automatically set a more appropriate scale. (Hint: you can make use of the
maxandminmethods to help.)Solution
# One method axes3.set_ylabel('min') axes3.plot(numpy.min(data, axis=0)) axes3.set_ylim(0,6)Solution
# A more automated approach min_data = numpy.min(data, axis=0) axes3.set_ylabel('min') axes3.plot(min_data) axes3.set_ylim(numpy.min(min_data), numpy.max(min_data) * 1.1)
Drawing Straight Lines
In the center and right subplots above, we expect all lines to look like step functions because non-integer value are not realistic for the minimum and maximum values. However, you can see that the lines are not always vertical or horizontal, and in particular the step function in the subplot on the right looks slanted. Why is this?
Solution
Because matplotlib interpolates (draws a straight line) between the points. One way to do avoid this is to use the Matplotlib
drawstyleoption:import numpy import matplotlib.pyplot data = numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',') fig = matplotlib.pyplot.figure(figsize=(10.0, 3.0)) axes1 = fig.add_subplot(1, 3, 1) axes2 = fig.add_subplot(1, 3, 2) axes3 = fig.add_subplot(1, 3, 3) axes1.set_ylabel('average') axes1.plot(numpy.mean(data, axis=0), drawstyle='steps-mid') axes2.set_ylabel('max') axes2.plot(numpy.max(data, axis=0), drawstyle='steps-mid') axes3.set_ylabel('min') axes3.plot(numpy.min(data, axis=0), drawstyle='steps-mid') fig.tight_layout() matplotlib.pyplot.show()
Make Your Own Plot
Create a plot showing the standard deviation (
numpy.std) of the inflammation data for each day across all patients.Solution
std_plot = matplotlib.pyplot.plot(numpy.std(data, axis=0)) matplotlib.pyplot.show()
Moving Plots Around
Modify the program to display the three plots on top of one another instead of side by side.
Solution
import numpy import matplotlib.pyplot data = numpy.loadtxt(fname='combined-no-headers.csv', delimiter=',') array = np.delete(array, 0, 1) # change figsize (swap width and height) fig = matplotlib.pyplot.figure(figsize=(3.0, 10.0)) # change add_subplot (swap first two parameters) axes1 = fig.add_subplot(3, 1, 1) axes2 = fig.add_subplot(3, 1, 2) axes3 = fig.add_subplot(3, 1, 3) axes1.set_ylabel('average') axes1.plot(numpy.mean(data, axis=0)) axes2.set_ylabel('max') axes2.plot(numpy.max(data, axis=0)) axes3.set_ylabel('min') axes3.plot(numpy.min(data, axis=0)) fig.tight_layout() matplotlib.pyplot.show()
Pandas
Pandas is a powerful dataframe tool that allows you to interact with data in a manner more like an Excel sheet or R - it’s the arrays we have been using in Numpy, but with labeling retained!
import pandas as pd
df = pd.read_csv('Acyclic.csv')
print(df)
Unnamed: 0 %top ER1 ER2 ER3 ER4 ER5 ER6 ER7 EZ ... \
0 0 2.25 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
1 1 2.50 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
2 2 5.00 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
3 3 5.00 -0.09 0.00 0.0 -0.09 0.0 0.0 0.00 1 ...
4 4 3.50 -0.01 -0.17 0.0 -0.01 0.0 0.0 -0.17 1 ...
.. ... ... ... ... ... ... ... ... ... .. ...
174 174 2.50 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
175 175 3.25 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
176 176 5.00 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
177 177 2.00 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
178 178 15.00 0.36 0.00 0.0 -0.17 0.0 0.0 -0.15 1 ...
boron reagent equivs boron reagent \
0 OB(C1=CC=CC=C1)O 2.0
1 OB(C1=CC=CC=C1)O 2.0
2 OB(C1=CC=CC=C1)O 3.0
3 C1(B2OB(C3=CC=CC=C3)OB(C4=CC=CC=C4)O2)=CC=CC=C1 2.0
4 OB(C1=CC=CC=C1)O 2.0
.. ... ...
174 OB(C1=CC=CC=C1)O 2.4
175 CC1=CC(B(O)O)=CC=C1 2.4
176 OB(C1=CC=CC=C1)O 2.0
177 OB(C1=CC=CC=C1)O 2.4
178 OB(C1=CC=CC=C1)O 2.4
ligand mol% Rh mol% ligand solvent \
0 L3.1 1.5 3.0 dioxane/H2O
1 L3.1 1.5 3.3 dioxane/H2O
2 L3.1 2.5 5.5 dioxane/H2O
3 L3.1 1.5 3.0 dioxane/H2O
4 L3.10 1.5 3.6 dioxane
.. ... ... ... ...
174 L3.86 2.5 6.0 dioxane/H2O
175 L3.86 2.5 6.0 dioxane/H2O
176 L3.86 2.5 6.0 dioxane/H2O
177 L3.86 2.5 6.0 dioxane/H2O
178 L3.87 2.5 6.0 dioxane/H2O
substrate temp time yield
0 CC(/C=C/C(C)C)=O 30 3.0 81.0
1 CC(/C=C/CCCCC)=O 30 1.0 84.0
2 O=C(/C=C/C(OC(C)(C)C)=O)OC(C)(C)C 50 3.0 78.0
3 O=C(/C=C/CCC)OC(C)C 50 1.0 73.0
4 CC(/C=C/C(C)C)=O 25 1.0 91.0
.. ... ... ... ...
174 CCCC/C=C/C1=NC2=CC=CC=C2C=C1 80 0.5 56.0
175 CCCC/C=C/C1=NC2=CC=CC=C2C=C1 80 0.5 71.0
176 C12=CC=CC=C1N=CC(/C=C/CCC3=CC=CC=C3)=N2 80 0.5 80.0
177 O=[N+](C1=CC=C(/C=C/CCC2=CC=CC=C2)C=C1)[O-] 80 0.5 35.0
178 O=[N+](C1=CC=C(/C=C/CCC2=CC=CC=C2)C=C1)[O-] 120 0.5 76.0
[179 rows x 40 columns]
This is dramatically different to the array function where we worked with a csv file where the headers had been stripped out, and where we stripped out the first column - the index. Printing this dataframe is very informative, displaying the headers and the indices, plus the size of dataframe. The best thing is we can now do some work using those labels!
There is a lot of data so let’s filter it and remove some columns.
df = df.filter(['solvent', '%top', 'temp', 'time', 'yield'])
print(df)
print(df['solvent']) # print just the solvent column
solvent %top temp time yield
0 dioxane/H2O 2.25 30 3.0 81.0
1 dioxane/H2O 2.50 30 1.0 84.0
2 dioxane/H2O 5.00 50 3.0 78.0
3 dioxane/H2O 5.00 50 1.0 73.0
4 dioxane 3.50 25 1.0 91.0
.. ... ... ... ... ...
174 dioxane/H2O 2.50 80 0.5 56.0
175 dioxane/H2O 3.25 80 0.5 71.0
176 dioxane/H2O 5.00 80 0.5 80.0
177 dioxane/H2O 2.00 80 0.5 35.0
178 dioxane/H2O 15.00 120 0.5 76.0
[179 rows x 5 columns]
0 dioxane/H2O
1 dioxane/H2O
2 dioxane/H2O
3 dioxane/H2O
4 dioxane
...
174 dioxane/H2O
175 dioxane/H2O
176 dioxane/H2O
177 dioxane/H2O
178 dioxane/H2O
Name: solvent, Length: 179, dtype: object
Printing a column now is much more human readable as we just need the name of the column. And when we want to do some work upon that data, we can convert first to a normal array.
df.to_numpy()
array([['dioxane/H2O', 2.25, 30, 3.0, 81.0],
['dioxane/H2O', 2.5, 30, 1.0, 84.0],
['dioxane/H2O', 5.0, 50, 3.0, 78.0],
['dioxane/H2O', 5.0, 50, 1.0, 73.0],
['dioxane', 3.5, 25, 1.0, 91.0],
['MeOH/CH2Cl2', 33.0, 60, 1.0, 89.0],
['MeOH/CH2Cl2', 11.5, 60, 1.0, 99.0],
['MeOH/CH2Cl2', 1.275, 60, 1.0, 92.0],
['i-PrOH/CH2Cl2', 1.833333333, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.666666667, 60, 1.0, 85.0],
['i-PrOH/CH2Cl2', 2.166666667, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.75, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.25, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 2.0, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.0, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.0, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 0.5, 60, 1.0, 99.0],
['i-PrOH/CH2Cl2', 1.5, 60, 1.0, 77.0],
['i-PrOH/CH2Cl2', 1.75, 60, 1.0, 99.0],
['MeOH/CH2Cl2', 1.0, 60, 1.0, 83.0],
['i-PrOH/CH2Cl2', 2.5, 60, 1.0, 74.0],
['toluene/H2O', 25.0, 50, 23.0, 67.0],
['dioxane', 11.5, 25, 1.0, 90.0],
['toluene/H2O', 23.0, 50, 23.0, 85.0],
['dioxane', 11.0, 25, 1.0, 72.0],
['dioxane', 8.5, 25, 1.0, 100.0],
['toluene/H2O', 22.5, 50, 23.0, 56.0],
['EtOH', 2.0, 30, 2.0, 89.0],
['EtOH', 2.0, 30, 2.0, 89.0],
['EtOH', 3.5, 30, 8.0, 99.0],
['toluene/H2O', 28.0, 50, 23.0, 98.0],
['dioxane', 4.5, 25, 1.0, 85.0],
['toluene/H2O', 27.5, 50, 23.0, 90.0],
['dioxane', 3.625, 25, 1.0, 87.0],
['EtOH', 1.0, 25, 96.0, 68.0],
['EtOH', 3.25, 25, 11.0, 86.0],
['EtOH', 1.5, 25, 5.5, 72.0],
['EtOH', 2.0, 25, 50.0, 74.0],
['EtOH', 2.5, 25, 25.0, 77.0],
['EtOH', 5.0, 25, 24.0, 94.0],
['EtOH', 1.4, 25, 96.0, 80.0],
['EtOH', 2.75, 25, 16.0, 99.0],
['EtOH', 0.3, 25, 72.0, 99.0],
['EtOH', 11.0, 25, 27.0, 98.0],
['EtOH', 3.0, 25, 16.0, 95.0],
['EtOH', 1.0, 60, 2.0, 81.0],
['EtOH', 0.5, 60, 14.0, 99.5],
['EtOH', 2.0, 60, 23.0, 88.0],
['toluene/H2O', 8.125, 50, 1.5, 85.0],
['dioxane/H2O', 11.0, 30, 1.0, 89.0],
['dioxane/H2O', 4.0, 30, 1.0, 90.0],
['dioxane', 6.0, 25, 1.0, 74.0],
['toluene/H2O', 24.0, 50, 23.0, 95.0],
['dioxane/H2O', 11.0, 30, 1.0, 91.0],
['dioxane/H2O', 7.5, 30, 1.0, 88.0],
['dioxane/H2O', 19.5, 25, 3.0, 95.0],
['MeOH/H2O', 5.0, 50, 1.0, 91.0],
['dioxane/H2O', 5.0, 50, 3.0, 90.0],
['dioxane/H2O', 4.5, 50, 3.0, 80.0],
['dioxane/H2O', 7.0, 50, 3.0, 78.0],
['dioxane/H2O', 5.0, 50, 3.0, 85.0],
['dioxane/H2O', 6.5, 50, 3.0, 91.0],
['dioxane', 1.5, 50, 6.0, 90.0],
['dioxane/H2O', 7.5, 30, 1.0, 95.0],
['dioxane/H2O', 5.75, 30, 1.0, 90.0],
['dioxane/H2O', 1.0, 30, 1.0, 90.0],
['dioxane/H2O', 2.0, 30, 3.0, 93.0],
['dioxane/H2O', 2.5, 30, 1.0, 93.0],
['dioxane', 21.5, 20, 1.0, 99.0],
['dioxane', 1.5, 20, 1.0, 95.0],
['dioxane', 2.0, 20, 1.0, 96.0],
['dioxane', 0.5, 20, 1.0, 94.0],
['dioxane', 1.0, 20, 1.0, 99.0],
['dioxane', 6.25, 20, 1.0, 33.0],
['dioxane', 1.75, 20, 1.0, 95.0],
['dioxane', 1.0, 20, 1.0, 92.0],
['dioxane', 2.0, 20, 1.0, 90.0],
['dioxane', 2.0, 20, 1.0, 92.0],
['dioxane', 1.0, 30, 2.0, 96.0],
['dioxane', 1.0, 30, 2.0, 97.0],
['dioxane', 1.0, 20, 1.0, 91.0],
['dioxane/H2O', 15.0, 50, 20.0, 75.0],
['dioxane', 24.0, 20, 1.0, 74.0],
['dioxane/H2O', 96.0, 25, 3.0, 99.0],
['dioxane/H2O', 91.5, 25, 3.0, 98.0],
['dioxane', 0.5, 50, 6.0, 95.0],
['dioxane', 0.5, 50, 6.0, 94.0],
['dioxane', 3.5, 50, 6.0, 90.0],
['dioxane', 2.5, 50, 6.0, 92.0],
['dioxane', 2.5, 50, 6.0, 94.0],
['dioxane', 2.0, 50, 6.0, 95.0],
['dioxane', 3.5, 50, 6.0, 96.0],
['dioxane', 1.5, 50, 12.0, 91.0],
['dioxane', 1.0, 50, 6.0, 89.0],
['dioxane/H2O', 4.2, 50, 6.0, 94.0],
['dioxane/H2O', 2.5, 50, 6.0, 88.0],
['dioxane/H2O', 95.0, 50, 20.0, 92.0],
['dioxane/H2O', 94.5, 50, 20.0, 83.0],
['dioxane/H2O', 93.5, 50, 20.0, 93.0],
['dioxane/H2O', 96.0, 50, 20.0, 84.0],
['dioxane/H2O', 94.5, 50, 20.0, 83.0],
['dioxane/H2O', 92.5, 50, 20.0, 92.0],
['dioxane/H2O', 96.0, 50, 20.0, 92.0],
['dioxane/H2O', 93.5, 50, 20.0, 86.0],
['dioxane/H2O', 93.0, 50, 20.0, 74.0],
['dioxane/H2O', 90.0, 50, 20.0, 91.0],
['DCM', 16.5, 25, 1.0, 99.0],
['MeOH/CH2Cl2', 24.0, 25, 1.0, 99.0],
['DCM', 2.5, 30, 3.0, 99.0],
['MeOH/CH2Cl2', 1.5, 30, 3.0, 99.0],
['dioxane/H2O', 25.5, 25, 3.0, 99.0],
['dioxane/H2O', 16.5, 25, 3.0, 98.0],
['dioxane/H2O', 21.5, 25, 3.0, 98.0],
['MeOH/CH2Cl2', 12.75, 30, 3.0, 95.0],
['MeOH/CH2Cl2', 13.5, 30, 3.0, 98.0],
['MeOH/CH2Cl2', 7.625, 30, 3.0, 93.0],
['MeOH/CH2Cl2', 12.5, 30, 3.0, 99.0],
['MeOH/CH2Cl2', 14.75, 30, 3.0, 96.0],
['MeOH/CH2Cl2', 6.25, 30, 3.0, 43.0],
['dioxane/H2O', 14.0, 25, 3.0, 98.0],
['dioxane/H2O', 15.0, 80, 16.0, 73.0],
['dioxane/H2O', 7.0, 25, 2.0, 74.0],
['MeOH/CH2Cl2', 95.0, 25, 1.0, 40.0],
['dioxane/H2O', 95.0, 25, 0.1, 68.0],
['dioxane/H2O', 94.5, 25, 1.0, 78.0],
['dioxane/H2O', 94.0, 25, 1.0, 93.0],
['dioxane/H2O', 4.5, 25, 3.0, 99.0],
['dioxane/H2O', 91.5, 50, 1.25, 50.0],
['MeOH/H2O', 96.5, 50, 1.25, 63.0],
['MeOH/H2O', 94.5, 50, 22.0, 70.0],
['MeOH/H2O', 95.5, 50, 2.5, 85.0],
['dioxane/H2O', 96.0, 50, 1.25, 43.0],
['MeOH/H2O', 96.5, 50, 1.25, 90.0],
['MeOH/H2O', 95.5, 50, 4.0, 76.0],
['MeOH/H2O', 96.5, 50, 1.25, 78.0],
['MeOH/H2O', 95.5, 50, 1.25, 87.0],
['MeOH/H2O', 95.0, 50, 1.25, 78.0],
['MeOH/H2O', 95.5, 50, 14.0, 68.0],
['MeOH/H2O', 96.0, 50, 14.0, 62.0],
['dioxane/H2O', 95.0, 50, 14.0, 65.0],
['dioxane/H2O', 96.5, 50, 14.0, 70.0],
['MeOH/H2O', 96.0, 50, 24.0, 69.0],
['MeOH/H2O', 96.0, 50, 1.25, 76.0],
['MeOH/H2O', 96.25, 50, 1.25, 85.0],
['MeOH/H2O', 96.0, 50, 1.25, 78.0],
['MeOH/H2O', 96.0, 50, 1.25, 84.0],
['MeOH/H2O', 97.0, 50, 1.5, 95.0],
['MeOH/H2O', 96.5, 50, 2.0, 93.0],
['MeOH/H2O', 95.5, 50, 18.5, 95.0],
['dioxane/H2O', 74.0, 50, 1.25, 43.0],
['dioxane/H2O', 59.5, 50, 18.0, 71.0],
['dioxane/H2O', 95.5, 50, 18.0, 85.0],
['dioxane/H2O', 80.0, 50, 1.25, 45.0],
['dioxane', 0.3, 50, 3.0, 97.0],
['dioxane/H2O', 3.6, 30, 3.0, 84.0],
['toluene/H2O', 97.5, 100, 12.0, 83.0],
['toluene/H2O', 96.0, 100, 12.0, 72.0],
['toluene/H2O', 97.0, 100, 18.0, 88.0],
['dioxane/H2O', 5.0, 80, 16.0, 41.0],
['toluene', 98.0, 100, 12.0, 82.0],
['dioxane/H2O', 6.5, 80, 0.5, 84.0],
['dioxane/H2O', 2.5, 80, 0.5, 24.0],
['dioxane/H2O', 5.0, 80, 0.5, 44.0],
['dioxane/H2O', 4.5, 80, 0.5, 91.0],
['dioxane/H2O', 2.5, 80, 16.0, 72.0],
['dioxane/H2O', 2.5, 100, 16.0, 62.0],
['dioxane/H2O', 4.0, 100, 16.0, 56.0],
['dioxane/H2O', 3.5, 80, 1.5, 88.0],
['dioxane/H2O', 6.5, 80, 1.0, 79.0],
['dioxane/H2O', 3.0, 80, 0.5, 95.0],
['dioxane/H2O', 1.0, 80, 0.5, 72.0],
['dioxane/H2O', 1.5, 80, 0.5, 65.0],
['dioxane/H2O', 3.5, 80, 0.5, 69.0],
['dioxane/H2O', 3.5, 80, 0.5, 78.0],
['dioxane/H2O', 2.5, 80, 0.5, 56.0],
['dioxane/H2O', 3.25, 80, 0.5, 71.0],
['dioxane/H2O', 5.0, 80, 0.5, 80.0],
['dioxane/H2O', 2.0, 80, 0.5, 35.0],
['dioxane/H2O', 15.0, 120, 0.5, 76.0]], dtype=object)
Key Points
Use the
pyplotmodule from thematplotliblibrary for creating simple visualizations.
